Shalaka Chitale, Ph.D.
Head of Biology & Data Science
Cell-based disease models are an essential tool for biomedical innovation, providing insights on mechanisms of pathology as well as cellular responses to therapeutics. In this interview, Dr. Chitale discusses the current state of in vitro disease modeling and how Pixel™ is allowing scientists to glimpse cell biology with never-before-seen detail and throughput.
Was there a specific application for which Pixel technology was initially developed?
The foundation for what would become Pixel technology essentially came from trying to use CMOS electrodes to gain intracellular access to neurons and measure their activity. From these early invasive measurements, we realized that we could use the electrodes to measure cells non-invasively, allowing us to observe them over time. This led to the development of electrical imaging: using impedance-based electrical field measurements to generate a wealth of morphological and functional information on any cell type.
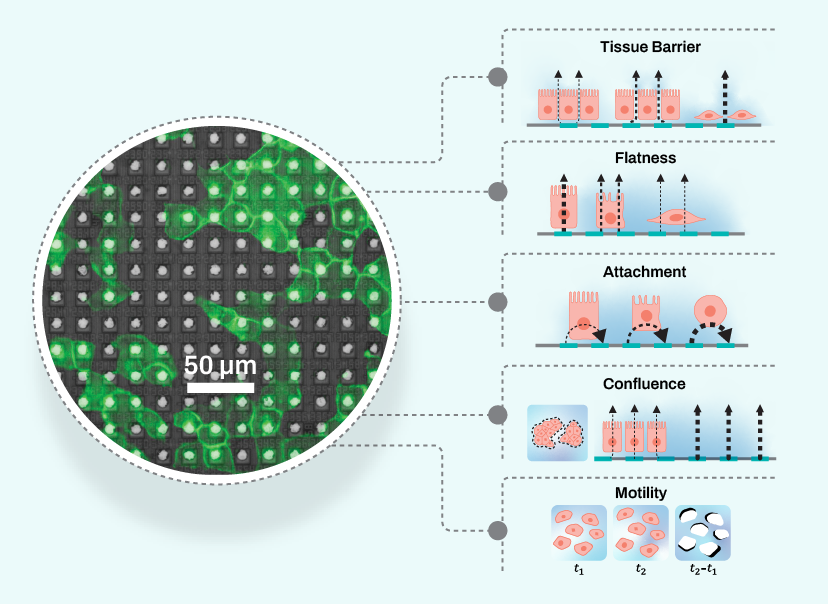
Impedance-based measurements can be used to quantify a variety of cellular properties.
How did new applications emerge?
Early on, we used electrical imaging to measure tissue barrier in endothelial and epithelial cell cultures. Prior to this, impedance readouts had been primarily used for similar barrier measurements. However, our technology stood out as being more accurate and sensitive than other methods.
We began to realize that we could measure many more parameters in many different cell types, and not just in monolayers but also in 3D. Eventually, we were able to integrate electrophysiological measurements with electrical imaging so that the same electrodes can be repurposed to measure the same cells in one plate. Increasing the density of the electrodes then enabled us to capture data at single-cell resolution.
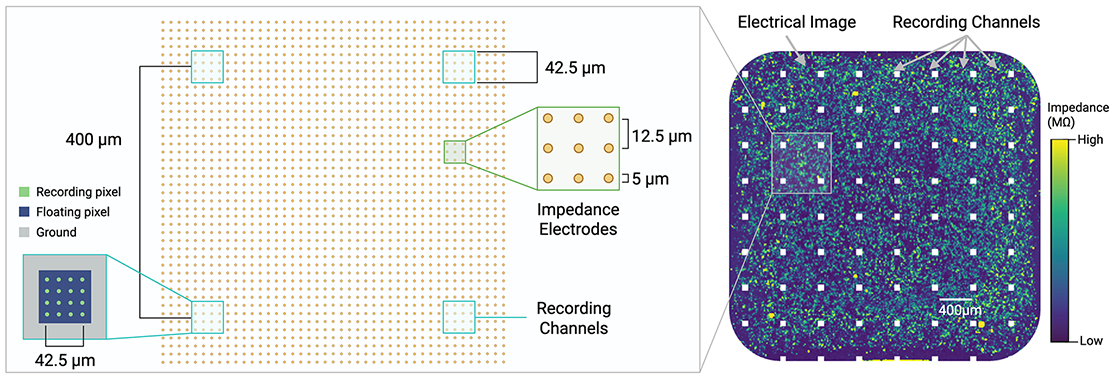
Pixel’s uniquely flexible microelectrode array repurposes electrodes to take different kinds of measurements.
How is Pixel changing how we study disease models?
There are many cellular changes that lead to disease states. The current approach for studying disease models is very linear: trying to address one cause/phenotype at a time in separate assays. With this approach, you’re not capturing the entire disease phenotype, which makes it difficult to come up with a comprehensive treatment or solution. This is one of the major reasons why something may work in an in vitro assay but not in an animal model.
With Pixel, you can truly see the entire disease phenotype at the same time. For example, a hallmark of some neurodegenerative diseases is the accumulation of certain proteins. These proteins not only change the structure of the neurons, but also their connectivity, how they fire and interact with each other. Pixel enables you to look at those changes simultaneously and see how your drug or treatment impacts all these factors.
Can you give a few examples of Pixel users deriving novel insights about their disease model?
One group was studying how genetic variants of iPSC-derived cardiomyocytes function differently. Pixel enabled them to compare these variants across multiple parameters like morphology, beat rate, and contractility using just one assay on the same plate of cells. With iPSCs, or induced pluripotent stem cells, you want to get as much information as possible from every cell, because it’s difficult and expensive to generate them. If they’re from patient-derived cells, that makes them even more precious.
Another group was investigating iPSC-derived fibroblasts in a neurodegenerative disease model. These were taken from patients with varying severity of the disease. The traditional readout for this kind of assay would be looking at the aggregation levels of a specific protein, which requires a lot of cells. Similarly to the cardiomyocyte assay, Pixel provided multiparametric data on how these fibroblasts differed from each other, all in the same assay. The cells reflected the relative severity of disease that was present in the patients.
The primary reason this group was using fibroblasts instead of neurons is because they’re easier to derive from iPSCs. But Pixel doesn’t require a lot of cells; with only one set of neurons, you can see the effects of protein aggregation across many cellular properties. Now that they’ve done this proof of principle experiment, they’re excited to move onto neuronal studies.
You mentioned that Pixel technology is uniquely adept at quantifying tissue barrier changes. What does tissue barrier have to do with disease modeling?
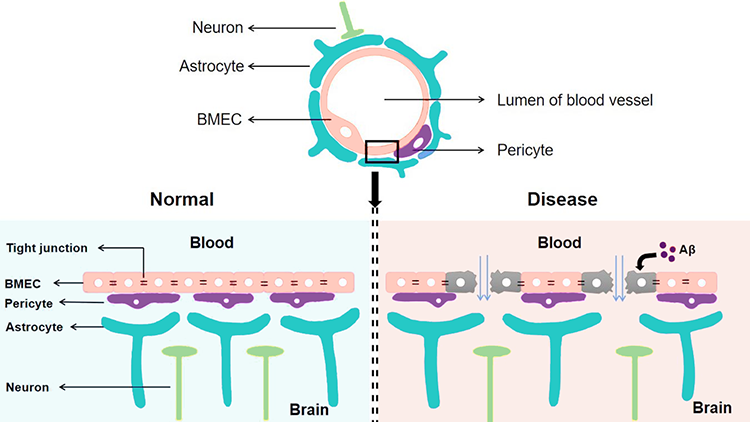
Tissue barriers are present all over the body – the blood-brain barrier, the lining of the lungs and the gut, and so on. Anywhere you need to keep two systems separated, disruption to these barriers can lead to disease, because things begin to interact that shouldn’t be. For example, “leakiness” in the blood-brain barrier can let in proteins and other inflammatory factors that affect the function of neurons and lead to neurodegenerative diseases.
Leakage in tissue barriers like the blood-brain barrier can lead to disease.
How does Pixel change how we study tissue barrier dysfunction?
The traditional way to measure tissue barrier is through a TEER assay, which essentially consists of putting one large electrode on either side of a cell sheet to measure how much of the current it blocks. But this method can be affected by imperfections in the cell sheet which may have nothing to do with the disease, just inconsistencies in how the cells grew. It’s also not very accurate because these measurements are being averaged over a large area, and the sensitivity is limited to one barrier type. You can’t tell whether the change in barrier is coming from cell death or from a loss of junction proteins.
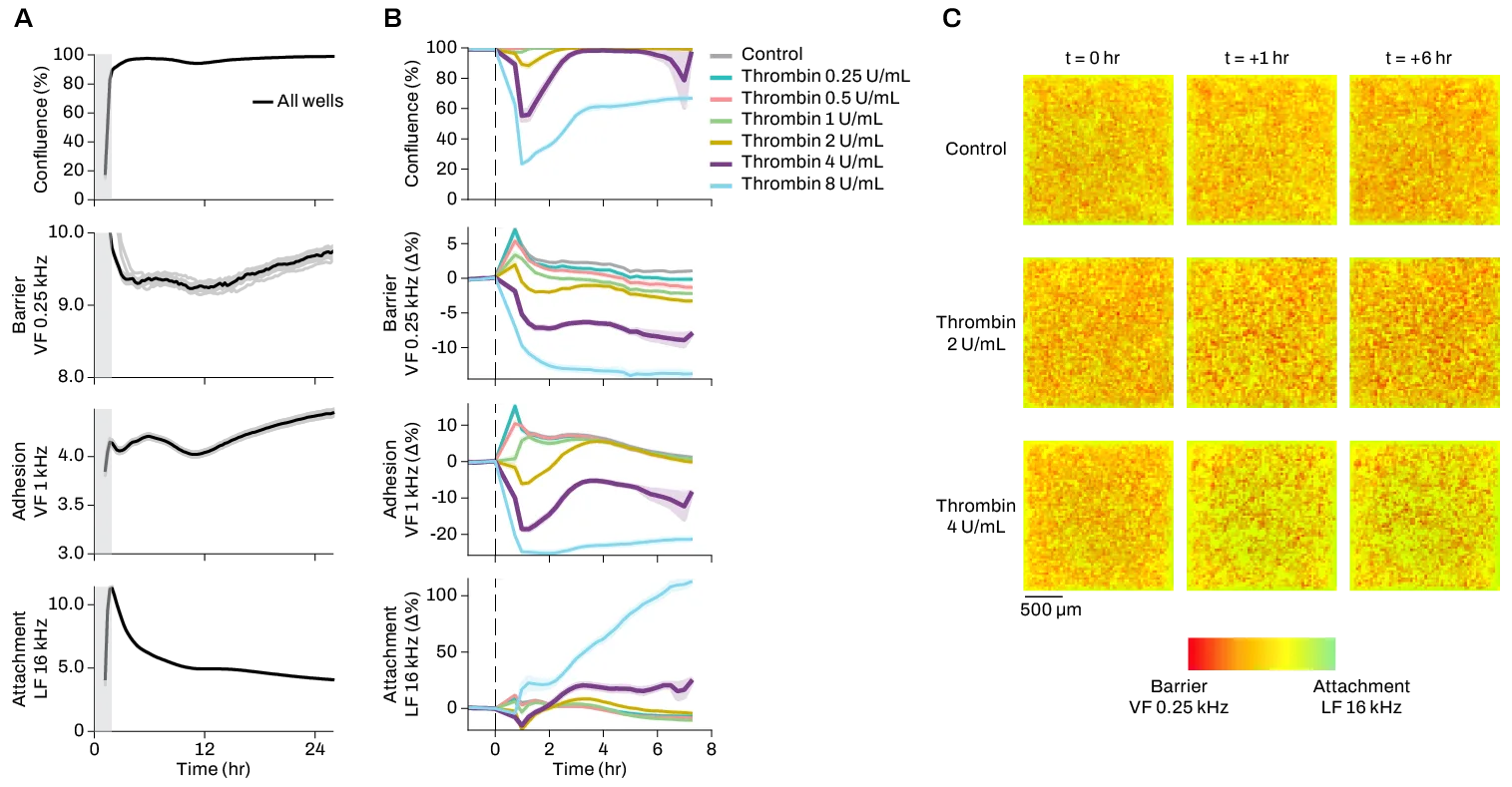
Pixel is much more accurate because it allows you to selectively look at intact areas of the cell sheet, leaving out regions with fewer or no cells. Its high spatial density of small, CMOS-based electrodes also makes it much more sensitive than large TEER electrodes. Additionally, because Pixel takes measurements at different frequences, we can study multiple types of barriers, like differentiating between adhesion junctions and tight junctions.
Many users have found that Pixel gives them more reproducible results, which enables them to use tissue barrier as a readout in high-throughput screens for the first time. Previously, you would screen for a proxy, like expression of a barrier-related protein, but that’s not the same thing as measuring actual formation of a barrier.
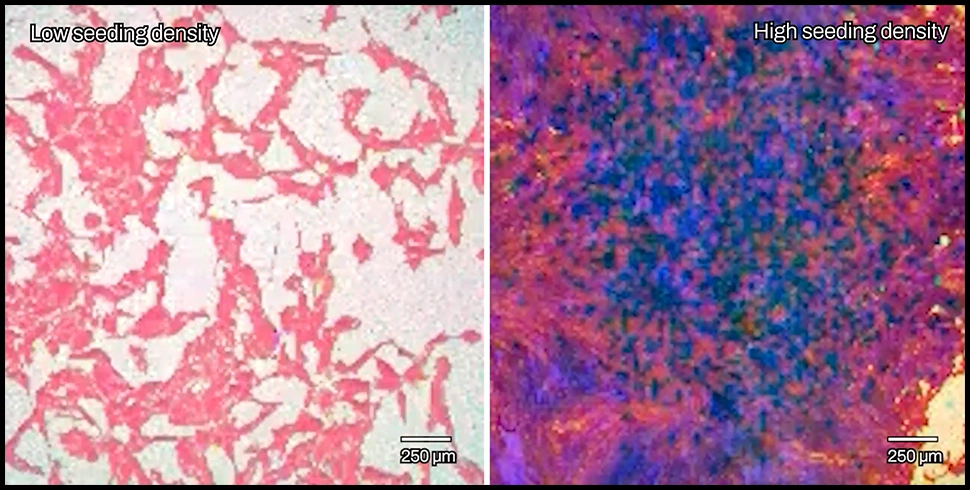
Researchers are moving towards complex cell models like 2D and 3D co-cultures to better understand heterogeneity in disease processes. How can Pixel help us understand these models?
The limitation of studying these cell models is always throughput. As your assay gets more biologically complex, you can either have a few readouts (basically, are cells alive or dead) in high throughput or many readouts in low throughput. That’s where Pixel comes in: it increases both the information density and the scale of the assay, which really matters when your cell model is expensive and difficult to grow.
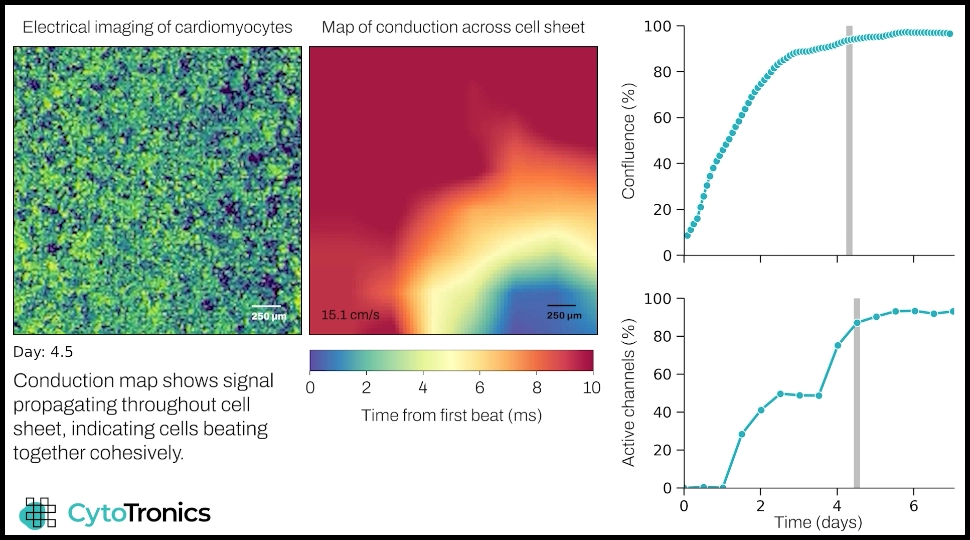
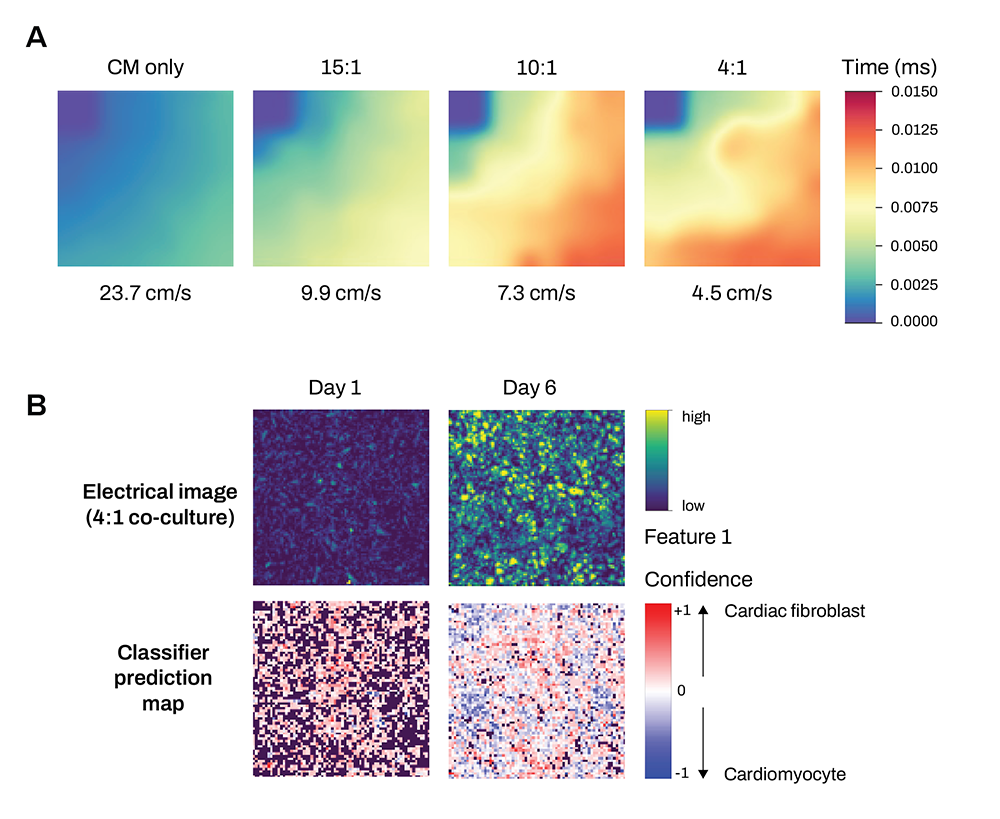
Electrical imaging combined with cardiac measurements can identify and correlate structural and functional changes in cardiac co-cultures relevant to disease models.
What have you seen users be most excited about?
I think seeing the electrophysiological data together with the electrical imaging data in the neural and cardiac applications has really been blowing people’s minds, and just how much you can learn about cell morphology with electrical imaging. When you look at these cells under a microscope, they don’t look very different, but electrical imaging reveals all these new layers of information that you couldn’t see by eye.
Another example that comes to mind is a user who was working with muscle cells. As these cells develop, they form these long tubes that bundle together to make up muscle fibers. They had seen all these steps optically before, but the electrical imaging data tracked them in such information-rich detail, and in a real-time, quantifiable way. People are used to looking at their cells at only a few timepoints, but being able to track them continuously opens entirely new avenues of exploration.